Simulation Notebook 6
Faces with JAX attractor network¶
This notebook demonstrates storage of face images in the FEP attractor network (JAX implementation), attractor dynamics, and reconstruction from noisy input.
Outline¶
Data: Load Olivetti faces (sklearn), downsample to keep runtime tractable, normalize to obtain patterns.
Training: Store face patterns in the network using the same local learning rule as in the scaling notebook.
Attractors: Evaluate deterministic retention (free run from stored pattern).
Reconstruction: Present noisy versions of stored faces as continuous bias; compare correlation of noisy input vs. reconstructed output with the original pattern.
Equations use the same notation as the manuscript: pattern , activations , coupling , and variational free energy (VFE) driving both inference and learning.
# Backend guard: force CPU (run before importing jax)
import os
os.environ["JAX_PLATFORMS"] = "cpu"
os.environ.pop("ENABLE_PJRT_COMPATIBILITY", None)
print("Configured JAX_PLATFORMS=cpu")Configured JAX_PLATFORMS=cpu
import numpy as np
import matplotlib.pyplot as plt
from scipy.ndimage import sobel
from sklearn.datasets import fetch_olivetti_faces
from simulation.network_jax import JAXAttractorNetwork1. Load and prepare face patterns¶
Olivetti faces: 400 images, 64×64 pixels. We downsample to 32×32 (1024 nodes) for tractable training. The network models binary evidence (for vs. against) at each node, so we preprocess images into binary edge maps: Sobel edge detection is applied, then gradient magnitude is thresholded to obtain patterns (edge = +1, no edge = −1). Faces remain recognizable from these edge maps. Flattened binary patterns are used for training and inference.
def grayscale_to_binary_edges(images_3d, edge_percentile=90):
"""
Convert (n, h, w) grayscale images to binary edge maps in {-1, +1}.
Edge pixels (high gradient magnitude) = +1, non-edge = -1.
Uses Sobel; threshold is per-image percentile so faces stay recognizable.
"""
n, h, w = images_3d.shape
out = np.empty((n, h, w), dtype=np.float64)
for i in range(n):
gx = sobel(images_3d[i], axis=1)
gy = sobel(images_3d[i], axis=0)
mag = np.sqrt(gx.astype(np.float64) ** 2 + gy.astype(np.float64) ** 2)
thresh = np.percentile(mag, edge_percentile)
out[i] = np.where(mag >= thresh, 1.0, -1.0)
return -1*out
def load_faces(subsample_size=64, n_max=np.inf, seed=0, binary_edges=False, edge_percentile=75):
"""Load Olivetti faces, downsample, optionally convert to binary edge maps; return (n, d) patterns."""
data = fetch_olivetti_faces()
images = data["images"] # (400, 64, 64)
step = 64 // subsample_size
small = images[:, ::step, ::step] # (n, h, w)
n, h, w = small.shape
n_keep = int(np.min((n, n_max)))
if n_keep < n:
rng = np.random.default_rng(seed)
idx = rng.choice(n, size=n_keep, replace=False)
small = small[idx]
n = n_keep
if binary_edges:
binary_3d = grayscale_to_binary_edges(small, edge_percentile=edge_percentile)
flat = binary_3d.reshape(n, -1).astype(np.float64) # values in {-1, +1}
flat = (flat - flat.mean(axis=1, keepdims=True)) / (flat.std(axis=1, keepdims=True) + 1e-8)
else:
flat = small.reshape(n, -1).astype(np.float64)
flat = (flat - flat.mean(axis=1, keepdims=True)) / (flat.std(axis=1, keepdims=True) + 1e-8)
return flat, (h, w)
patterns, shape = load_faces(n_max=np.inf, binary_edges=False, edge_percentile=75)
n_patterns, n_nodes = patterns.shape
print(f"Patterns: {n_patterns}, nodes: {n_nodes}, shape: {shape} (binary edge maps in {{-1, +1}})")Patterns: 400, nodes: 4096, shape: (64, 64) (binary edge maps in {-1, +1})
# Show a few stored faces (random selection)
rng_faces = np.random.default_rng(42)
idx_show = rng_faces.choice(len(patterns), size=10, replace=False)
fig, axes = plt.subplots(2, 5, figsize=(10, 4))
# Binary patterns in {-1, +1}; use fixed range for consistent contrast
for i, ax in enumerate(axes.flat):
ax.imshow(patterns[idx_show[i]].reshape(shape), cmap="gray", vmin=-1, vmax=1)
ax.set_axis_off()
plt.suptitle("Example stored face patterns")
plt.tight_layout(pad=0.02, w_pad=0.02, h_pad=0.02)
plt.show()
2. Train network to store faces¶
Same training protocol as in the scaling notebook: random pattern presentation, evidence level scales the input, local VFE-based updates.
cfg = {
"evidence_level": 1.0,
"beta": 0.7, #0.5
"lr": 0.0001,
"epochs": 10000,
"steps": 10,
"seed": 42,
}
jnet = JAXAttractorNetwork(n_nodes=n_nodes, seed=cfg["seed"])
vfe_hist = jnet.train(
patterns,
evidence_level=cfg["evidence_level"],
beta=cfg["beta"],
lr=cfg["lr"],
epochs=cfg["epochs"],
steps=cfg["steps"],
)
plt.figure(figsize=(6, 2))
plt.plot(np.asarray(vfe_hist), alpha=0.8)
plt.xlabel("Training step")
plt.ylabel("VFE")
plt.title("Training VFE")
plt.tight_layout(pad=0.02, w_pad=0.02, h_pad=0.02)
plt.show()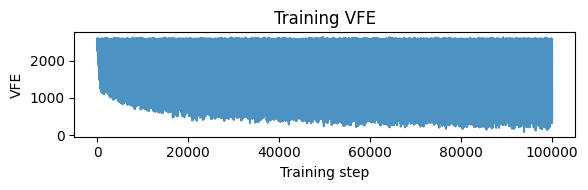
3. Attractor retention¶
Start from each stored pattern and run deterministic inference with no external bias (). The final state should remain close to the pattern if it lies near an attractor.
def _safe_corr(a, b):
c = np.corrcoef(a, b)[0, 1]
return 0.0 if np.isnan(c) else float(c)
def retention_jax(jnet, patterns, infer_beta=10.0, steps=150):
"""Mean correlation between relaxed state and original pattern (free run, no bias)."""
n = patterns.shape[1]
scores = []
for p in patterns:
acts, _ = jnet.infer(x0=p, u=np.zeros(n), beta=infer_beta, steps=steps, stochastic=False)
out = np.asarray(acts[-1])
scores.append(_safe_corr(out, p))
return float(np.mean(scores))
retention = retention_jax(jnet, patterns, infer_beta=100.0, steps=150)
print(f"Mean retention correlation (stored pattern vs attractor): {retention:.4f}")Mean retention correlation (stored pattern vs attractor): 0.4658
Visualize attractors¶
For a few stored patterns, run deterministic free dynamics () and show the stored pattern vs the attractor (final state). If the pattern is stored as an attractor, the two should match closely.
n_show_att = 6
rng_att = np.random.default_rng(43)
idx_att = rng_att.choice(len(patterns), size=n_show_att, replace=False)
fig, axes = plt.subplots(2, n_show_att, figsize=(2 * n_show_att, 4))
for j in range(n_show_att):
p = patterns[idx_att[j]]
acts, _ = jnet.infer(x0=p, u=np.zeros(n_nodes), beta=1, steps=150, stochastic=False)
attractor = np.asarray(acts[-1])
axes[0, j].imshow(p.reshape(shape), cmap="gray", vmin=-1, vmax=1)
axes[0, j].set_axis_off()
axes[1, j].imshow(attractor.reshape(shape), cmap="gray", vmin=-1, vmax=1)
axes[1, j].set_axis_off()
corr = _safe_corr(attractor, p)
axes[1, j].set_title(f"r={corr:.2f}")
axes[0, 0].set_ylabel("Stored pattern")
axes[1, 0].set_ylabel("Attractor")
plt.suptitle("Stored patterns vs fixed-point attractors (free run)")
plt.tight_layout(pad=0.02, w_pad=0.02, h_pad=0.02)
plt.show()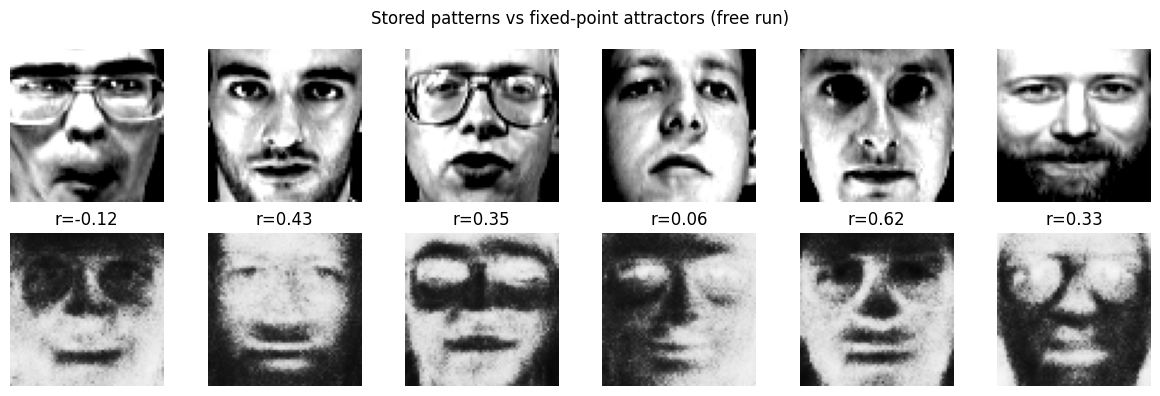
Alternative retention: random restarts and unique attractors¶
Another way to assess attractors: initialize with noise, run deterministic relaxation until convergence (change below a tolerance), repeat many times, then keep only unique fixed points (within a tolerance). Plot the unique attractors.
def relax_until_converged(jnet, x0, u, beta, tol=1e-4, max_steps=500):
"""Deterministic relaxation until max|x_new - x_prev| < tol or max_steps reached."""
x = np.asarray(x0, dtype=jnet.W.dtype)
u = np.asarray(u, dtype=jnet.W.dtype) if u is not None else np.zeros_like(x)
for _ in range(max_steps):
acts, _ = jnet.infer(x0=x, u=u, beta=beta, steps=1, stochastic=False)
x_new = np.asarray(acts[-1])
if np.max(np.abs(x_new - x)) < tol:
return x_new
x = x_new
return x_new
def unique_attractors(attractor_list, tol_unique=0.05):
"""Deduplicate attractors: two are the same if max|a - b| < tol_unique."""
unique = []
for a in attractor_list:
a = np.asarray(a).ravel()
is_new = True
for b in unique:
if np.max(np.abs(np.asarray(b).ravel() - a)) < tol_unique:
is_new = False
break
if is_new:
unique.append(a)
return unique
# Parameters
n_restarts = 200
infer_beta = 1.0
conv_tol = 1e-5
conv_max_steps = 500
tol_unique = 0.2
rng_restart = np.random.default_rng(44)
# Random initial states in [-1, 1], then relax to convergence
raw_attractors = []
for _ in range(n_restarts):
x0 = rng_restart.uniform(-1, 1, size=n_nodes).astype(np.float32)
x_final = relax_until_converged(jnet, x0, np.zeros(n_nodes), infer_beta, tol=conv_tol, max_steps=conv_max_steps)
raw_attractors.append(x_final)
unique = unique_attractors(raw_attractors, tol_unique=tol_unique)
print(f"Restarts: {n_restarts} → {len(unique)} unique attractors (tol={tol_unique})")
# Plot unique attractors (up to 15 in a grid)
n_show = min(30, len(unique))
n_col = 5
n_row = (n_show + n_col - 1) // n_col
fig, axes = plt.subplots(n_row, n_col, figsize=(2 * n_col, 2 * n_row))
axes = axes.ravel()
for i in range(n_show):
axes[i].imshow(unique[i].reshape(shape), cmap="gray", vmin=-1, vmax=1)
axes[i].set_axis_off()
axes[i].set_title(f"Attractor {i+1}")
for i in range(n_show, len(axes)):
axes[i].set_axis_off()
plt.suptitle("Unique attractors (noise init → deterministic relaxation)")
plt.tight_layout(pad=0.02, w_pad=0.02, h_pad=0.02)
plt.show()Restarts: 200 → 52 unique attractors (tol=0.2)

4. Noisy reconstruction¶
For each stored pattern, add Gaussian noise to form a noisy cue. Run stochastic inference with the noisy pattern as continuous bias . The mean activity after burn-in is the reconstruction; we expect correlation with the original to be higher than the correlation of the noisy input with the original (reconstruction gain).
def noisy_reconstruction_jax(jnet, patterns, noise_std=2.0, infer_beta=1.0, recon_steps=25, burn_in=5, repeats=1, seed=0):
"""Noisy input as bias; report mean noisy correlation and mean reconstructed correlation with original."""
rng = np.random.default_rng(seed)
n = patterns.shape[1]
noisy_corrs, recon_corrs = [], []
start_idx = min(max(0, burn_in), max(0, recon_steps - 1))
for p in patterns:
for _ in range(repeats):
noisy = p + rng.normal(0.0, noise_std * np.std(p), size=p.shape)
noisy_corrs.append(_safe_corr(noisy, p))
acts, _ = jnet.infer(x0=noisy, u=noisy, beta=infer_beta, steps=recon_steps, stochastic=True)
acts = np.asarray(acts)
mean_pattern = acts[start_idx:].mean(axis=0)
recon_corrs.append(_safe_corr(mean_pattern, p))
return float(np.mean(noisy_corrs)), float(np.mean(recon_corrs))
noisy_corr, recon_corr = noisy_reconstruction_jax(
jnet, patterns, noise_std=2.0, infer_beta=2.0, recon_steps=100, burn_in=5, seed=cfg["seed"] + 1000
)
gain = recon_corr - noisy_corr
print(f"Noisy input vs original: corr = {noisy_corr:.4f}")
print(f"Reconstructed vs original: corr = {recon_corr:.4f}")
print(f"Reconstruction gain: {gain:.4f}")Noisy input vs original: corr = 0.4479
Reconstructed vs original: corr = 0.7580
Reconstruction gain: 0.3100
5. Visualize reconstruction¶
For a few faces: show original, noisy input, and reconstructed (mean activity under noisy bias).
def reconstruct_one(jnet, original, noise_std=2.0, signal_strength=0.5, infer_beta=1.0, steps=500, burn_in=10, seed=0):
rng = np.random.default_rng(seed)
noisy = original + rng.normal(0.0, noise_std * np.std(original), size=original.shape)
noisy *= signal_strength
acts, _ = jnet.infer(x0=noisy, u=noisy, beta=infer_beta, steps=steps, stochastic=True)
acts = np.asarray(acts)
recon = acts[burn_in:].mean(axis=0)
return noisy, recon
n_show = 5
rng_recon = np.random.default_rng(cfg["seed"] * 42)
idx_recon = rng_recon.choice(len(patterns), size=n_show, replace=False)
fig, axes = plt.subplots(5, n_show, figsize=(1.5 * n_show, 9))
for j in range(n_show):
idx = idx_recon[j]
orig = patterns[idx]
axes[0, j].imshow(orig.reshape(shape), cmap="gray")
axes[0, j].set_axis_off()
#axes[0, j].set_title(f"Original") if j == 0 else ""
##################
infer_beta = 0.4
signal_strength = 2.0
noisy, recon = reconstruct_one(jnet, orig, noise_std=0.5, signal_strength=signal_strength, infer_beta=infer_beta, seed=int(rng_recon.integers(0, 1e6)))
axes[1, j].imshow(noisy.reshape(shape), cmap="gray")
#axes[1, j].set_title(f"Noisy input") if j == 0 else ""
axes[1, j].set_axis_off()
axes[2, j].imshow(recon.reshape(shape), cmap="gray")
#axes[2, j].set_title(f"Reconstructed (prior precision: {infer_beta} likelihood precision: {signal_strength})", loc='left') if j == 0 else ""
axes[2, j].set_axis_off()
##################
infer_beta = 0.7
signal_strength = 1.0
noisy, recon = reconstruct_one(jnet, orig, noise_std=0.5, signal_strength=signal_strength, infer_beta=infer_beta, seed=int(rng_recon.integers(0, 1e6)))
axes[3, j].imshow(recon.reshape(shape), cmap="gray")
#axes[3, j].set_title(f"Reconstructed (prior precision: {infer_beta} likelihood precision: {signal_strength})", loc='left') if j == 0 else ""
axes[3, j].set_axis_off()
##################
infer_beta = 1.0
signal_strength = 0.5
noisy, recon = reconstruct_one(jnet, orig, noise_std=0.5, signal_strength=signal_strength, infer_beta=infer_beta, seed=int(rng_recon.integers(0, 1e6)))
axes[4, j].imshow(recon.reshape(shape), cmap="gray")
#axes[4, j].set_title(f"Reconstructed (prior precision: {infer_beta} likelihood precision: {signal_strength})", loc='left') if j == 0 else ""
axes[4, j].set_axis_off()
axes[0, 2].set_title("Original")
axes[1, 2].set_title("Noisy input")
axes[2, 2].set_title(f"Reconstructed (prior precision: 0.4 likelihood precision: 0.9)")
axes[3, 2].set_title(f"Reconstructed (prior precision: 0.7 likelihood precision: 0.6)")
axes[4, 2].set_title(f"Reconstructed (prior precision: 1.0 likelihood precision: 0.3)")
plt.suptitle("Reconstruction from noisy face input")
plt.tight_layout(pad=0.02, w_pad=0.02, h_pad=0.02)
plt.show()
n_show = 5
rng_recon = np.random.default_rng(cfg["seed"] * 42)
idx_recon = rng_recon.choice(len(patterns), size=n_show, replace=False)
fig, axes = plt.subplots(5, n_show, figsize=(1.5 * n_show, 9))
for j in range(n_show):
idx = idx_recon[j]
orig = patterns[idx]
axes[0, j].imshow(orig.reshape(shape), cmap="gray")
axes[0, j].set_axis_off()
#axes[0, j].set_title(f"Original") if j == 0 else ""
##################
infer_beta = 0.4
signal_strength = 0.9
noisy, recon = reconstruct_one(jnet, orig, noise_std=2.0, signal_strength=signal_strength, infer_beta=infer_beta, seed=int(rng_recon.integers(0, 1e6)))
axes[1, j].imshow(noisy.reshape(shape), cmap="gray")
#axes[1, j].set_title(f"Noisy input") if j == 0 else ""
axes[1, j].set_axis_off()
axes[2, j].imshow(recon.reshape(shape), cmap="gray")
#axes[2, j].set_title(f"Reconstructed (prior precision: {infer_beta} likelihood precision: {signal_strength})", loc='left') if j == 0 else ""
axes[2, j].set_axis_off()
##################
infer_beta = 0.7
signal_strength = 0.6
noisy, recon = reconstruct_one(jnet, orig, noise_std=2.0, signal_strength=signal_strength, infer_beta=infer_beta, seed=int(rng_recon.integers(0, 1e6)))
axes[3, j].imshow(recon.reshape(shape), cmap="gray")
#axes[3, j].set_title(f"Reconstructed (prior precision: {infer_beta} likelihood precision: {signal_strength})", loc='left') if j == 0 else ""
axes[3, j].set_axis_off()
##################
infer_beta = 1.0
signal_strength = 0.3
noisy, recon = reconstruct_one(jnet, orig, noise_std=2.0, signal_strength=signal_strength, infer_beta=infer_beta, seed=int(rng_recon.integers(0, 1e6)))
axes[4, j].imshow(recon.reshape(shape), cmap="gray")
#axes[4, j].set_title(f"Reconstructed (prior precision: {infer_beta} likelihood precision: {signal_strength})", loc='left') if j == 0 else ""
axes[4, j].set_axis_off()
axes[0, 2].set_title("Original")
axes[1, 2].set_title("Noisy input")
axes[2, 2].set_title(f"Reconstructed (prior precision: 0.4 likelihood precision: 0.9)")
axes[3, 2].set_title(f"Reconstructed (prior precision: 0.7 likelihood precision: 0.6)")
axes[4, 2].set_title(f"Reconstructed (prior precision: 1.0 likelihood precision: 0.3)")
plt.suptitle("Reconstruction from noisy face input")
plt.tight_layout(pad=0.02, w_pad=0.02, h_pad=0.02)
plt.show()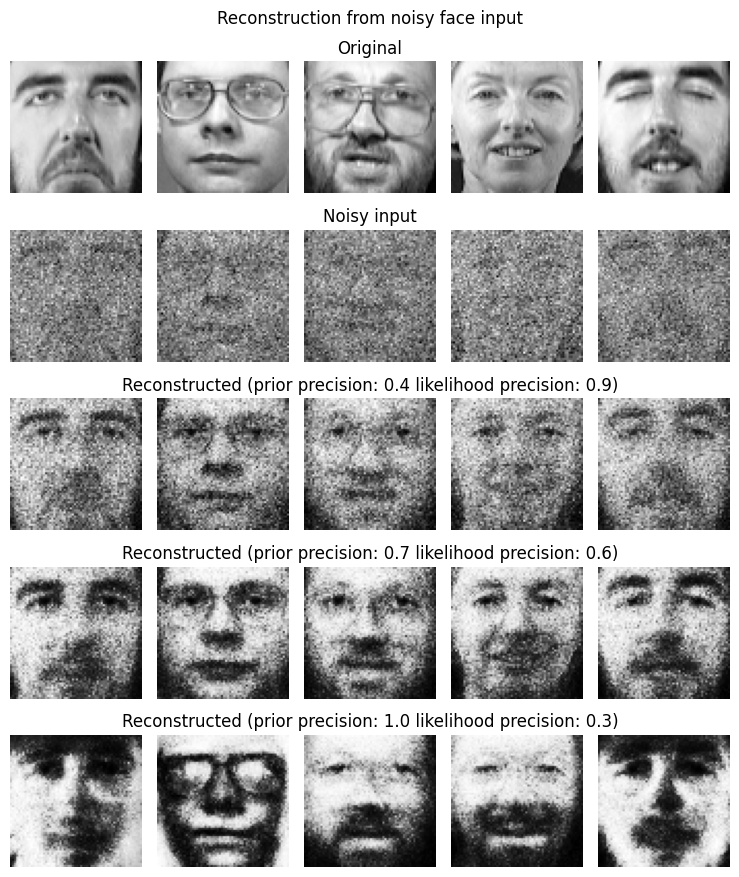
# 5x5 plot: row 0 = [empty, input, empty, noisy, empty]; rows 1-4 = 4x5 grid (rows = input precision, cols = prior precision)
rng_6x5 = np.random.default_rng(cfg["seed"] + 42)
idx_one = rng_6x5.choice(len(patterns))
orig_one = patterns[idx_one]
noise_std_one = 2.0
signal_for_noisy = 1.0
noisy_one, _ = reconstruct_one(
jnet, orig_one, noise_std=noise_std_one, signal_strength=signal_for_noisy,
infer_beta=0.6, seed=int(rng_6x5.integers(0, 1e6))
)
signal_strengths = [0.1, 0.2, 0.3, 0.4] # input (likelihood) precision → rows
prior_betas = [0.4, 0.6, 0.8, 1.0, 1.2] # prior precision → columns
recons_4x5 = np.zeros((4, 5, orig_one.size))
for i, sig in enumerate(signal_strengths):
for j, infer_beta in enumerate(prior_betas):
_, recon = reconstruct_one(
jnet, orig_one, noise_std=noise_std_one, signal_strength=sig,
infer_beta=infer_beta, steps=100, burn_in=10, seed=int(rng_6x5.integers(0, 1e6))
)
recons_4x5[i, j] = recon
fig, axes = plt.subplots(5, 5, figsize=(2 * 5, 2 * 5))
for c in range(5):
axes[0, c].set_axis_off()
axes[0, 1].imshow(orig_one.reshape(shape), cmap="gray", vmin=-1, vmax=1)
axes[0, 1].set_axis_off()
axes[0, 3].imshow(noisy_one.reshape(shape), cmap="gray")
axes[0, 3].set_axis_off()
for i in range(4):
for j in range(5):
axes[i + 1, j].imshow(recons_4x5[i, j].reshape(shape), cmap="gray", vmin=-1, vmax=1)
axes[i + 1, j].set_axis_off()
axes[0, 1].set_title("Input")
axes[0, 3].set_title("Noisy input")
plt.suptitle("Reconstruction: prior vs likelihood precision (one random face)")
fig.subplots_adjust(left=0.01, right=0.99, top=0.94, bottom=0.01, wspace=0.02, hspace=0.02)
plt.show()
6. Summary¶
Retention: Stored face patterns are fixed points (attractors) of the deterministic dynamics; retention correlation summarizes how well the network holds each pattern under free run.
Reconstruction: When the network receives a noisy version of a stored face as continuous bias, stochastic inference pulls activity toward the corresponding attractor; the reconstructed image (mean activity) typically correlates better with the original than the noisy input does (positive reconstruction gain).